Function for rendering plot displaying the mean time spent in each state of
a state sequence object using ggplot2
(Wickham 2016)
instead of base R's
plot function that is used by
TraMineR::seqplot
(Gabadinho et al. 2011)
.
Usage
ggseqmtplot(
seqdata,
no.n = FALSE,
group = NULL,
weighted = TRUE,
with.missing = FALSE,
border = FALSE,
error.bar = NULL,
error.caption = TRUE,
facet_scale = "fixed",
facet_ncol = NULL,
facet_nrow = NULL
)Arguments
- seqdata
State sequence object (class
stslist) created with theTraMineR::seqdeffunction.- no.n
specifies if number of (weighted) sequences is shown (default is
TRUE)- group
A vector of the same length as the sequence data indicating group membership. When not NULL, a distinct plot is generated for each level of group.
- weighted
Controls if weights (specified in
TraMineR::seqdef) should be used. Default isTRUE, i.e. if available weights are used- with.missing
Specifies if missing states should be considered when computing the state distributions (default is
FALSE).- border
if
TRUEbars are plotted with black outline; default isFALSE(also acceptsNULL)- error.bar
allows to add error bars either using the standard deviation
"SD"or the standard error"SE"; default plot is without error bars- error.caption
a caption is added if error bars are displayed; this default behavior can be turned off by setting the argument to
"FALSE"- facet_scale
Specifies if y-scale in faceted plot should be
"fixed"(default) or"free_y"- facet_ncol
Number of columns in faceted (i.e. grouped) plot
- facet_nrow
Number of rows in faceted (i.e. grouped) plot
Value
A mean time plot created by using ggplot2.
If stored as object the resulting list object (of class gg and ggplot) also
contains the data used for rendering the plot
Details
The information on time spent in different states is obtained by an
internal call of TraMineR::seqmeant. This
requires that the input data (seqdata) are stored as state sequence
object (class stslist) created with the
TraMineR::seqdef function. The resulting
output then is prepared to be plotted with
ggplot2::geom_bar. The data and
specifications used for rendering the plot can be obtained by storing the
plot as an object. The appearance of the plot can be adjusted just like with
every other ggplot (e.g., by changing the theme or the scale using +
and the respective functions).
References
Gabadinho A, Ritschard G, Müller NS, Studer M (2011).
“Analyzing and Visualizing State Sequences in R with TraMineR.”
Journal of Statistical Software, 40(4), 1–37.
doi:10.18637/jss.v040.i04
.
Wickham H (2016).
ggplot2: Elegant Graphics for Data Analysis, Use R!, 2nd ed. edition.
Springer, Cham.
doi:10.1007/978-3-319-24277-4
.
Examples
library(TraMineR)
library(ggplot2)
# Use example data from TraMineR: actcal data set
data(actcal)
# We use only a sample of 300 cases
set.seed(1)
actcal <- actcal[sample(nrow(actcal), 300), ]
actcal.lab <- c("> 37 hours", "19-36 hours", "1-18 hours", "no work")
actcal.seq <- seqdef(actcal, 13:24, labels = actcal.lab)
#> [>] 4 distinct states appear in the data:
#> 1 = A
#> 2 = B
#> 3 = C
#> 4 = D
#> [>] state coding:
#> [alphabet] [label] [long label]
#> 1 A A > 37 hours
#> 2 B B 19-36 hours
#> 3 C C 1-18 hours
#> 4 D D no work
#> [>] 300 sequences in the data set
#> [>] min/max sequence length: 12/12
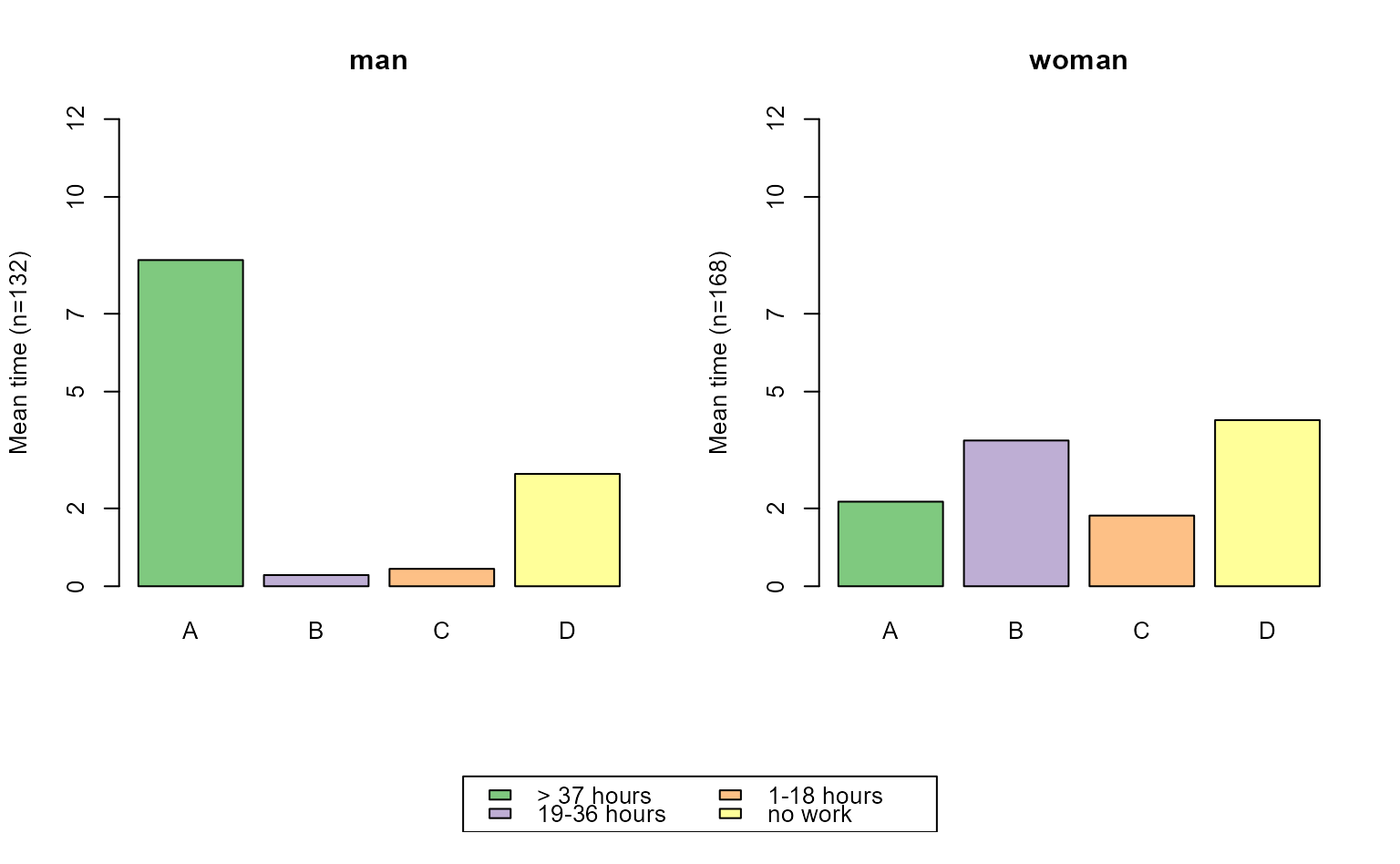
# modal state sequence plot; grouped by sex
# with TraMineR::seqplot
seqmtplot(actcal.seq, group = actcal$sex)
 # with ggseqplot
ggseqmtplot(actcal.seq, group = actcal$sex)
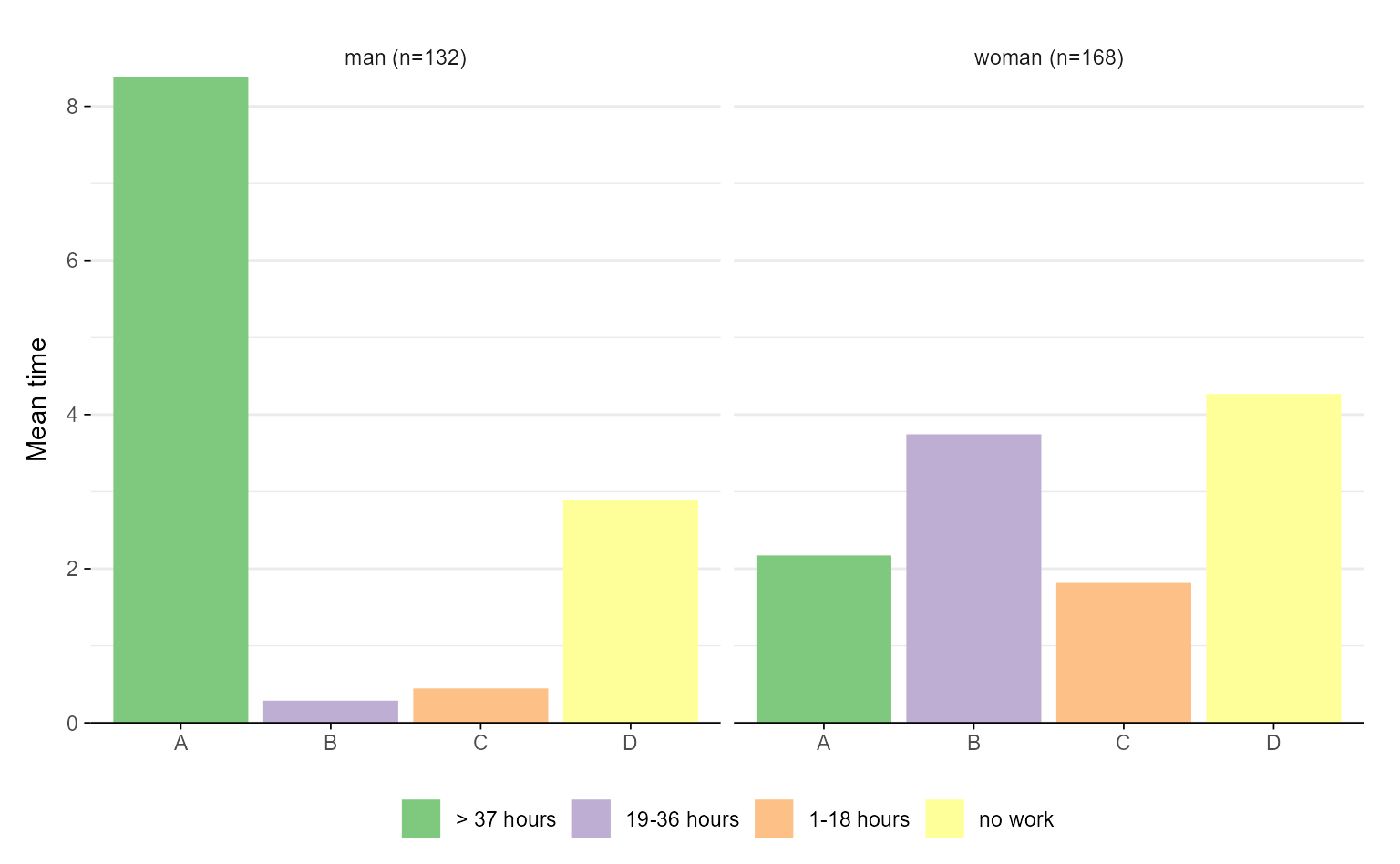
# with ggseqplot
ggseqmtplot(actcal.seq, group = actcal$sex)
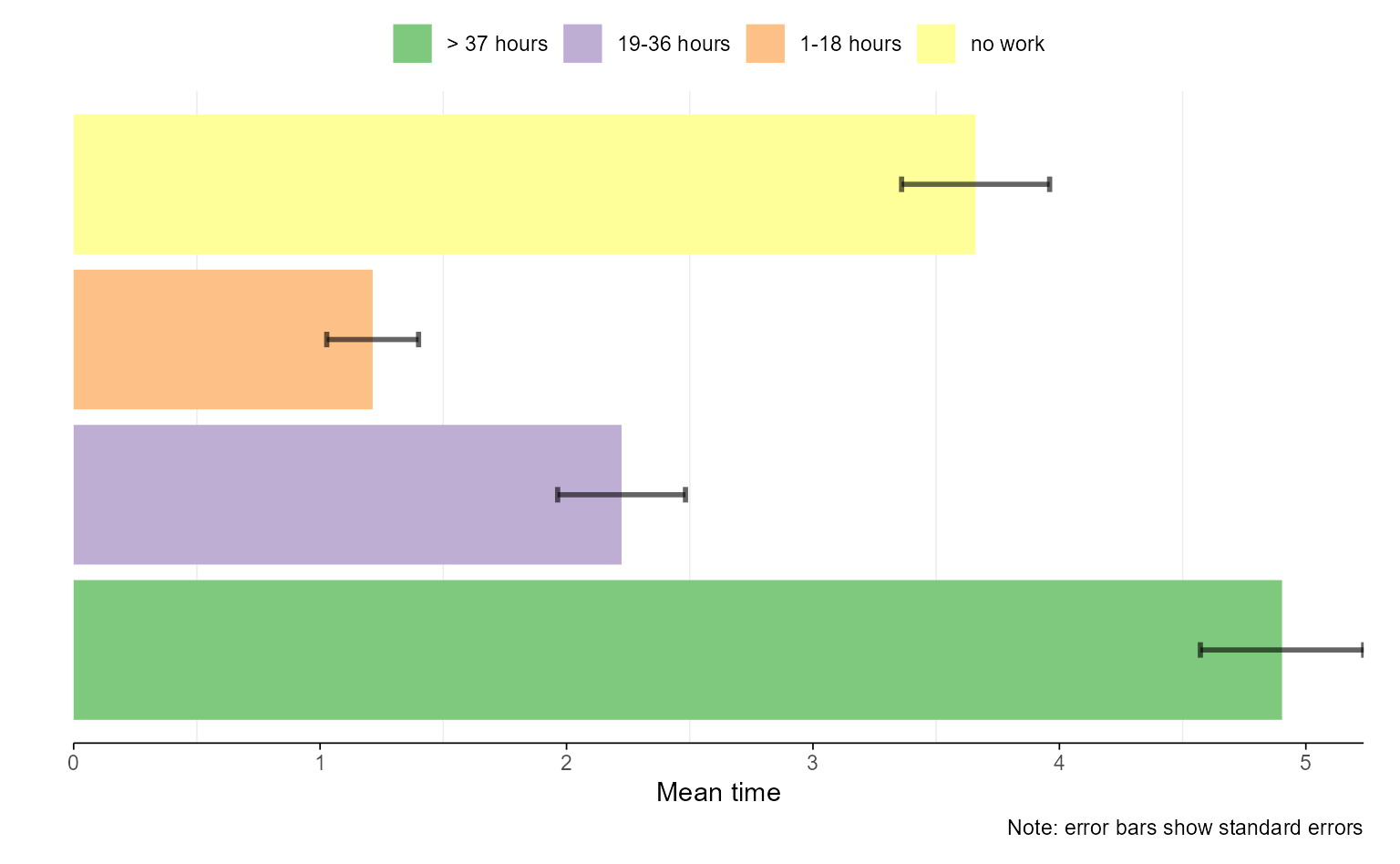
 # with ggseqplot using additional arguments and some adjustments
ggseqmtplot(actcal.seq, no.n = TRUE, error.bar = "SE") +
coord_flip() +
theme(axis.text.y=element_blank(),
axis.ticks.y = element_blank(),
panel.grid.major.y = element_blank(),
legend.position = "top")
# with ggseqplot using additional arguments and some adjustments
ggseqmtplot(actcal.seq, no.n = TRUE, error.bar = "SE") +
coord_flip() +
theme(axis.text.y=element_blank(),
axis.ticks.y = element_blank(),
panel.grid.major.y = element_blank(),
legend.position = "top")